New features in ver. 1.6
- Resouce databases are updated in human and Swiss-Prot dataset.
- PDB data for 3D structure assignment is updated in all dataset.
- Expression data from CAGE analyses are added to transcript cluster view.
- Some user interfaces are updated.
About this database
Alternative splicing (AS) is a molecular mechanism that produces multiple proteins from a single gene, and is thought to increase the number of proteins extensively over the number of genes in a genome, which could be required to generate complexity of higher eukaryote including human. However, what roles most of AS variants play are not known.
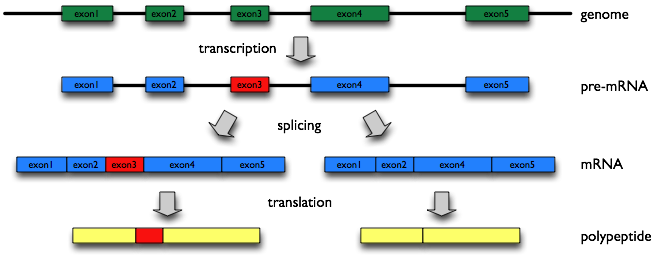
This database, AS-ALPS (Alternative Splicing-induced ALteration of Protein Structure), is aimed at providing useful information to analyze effect of AS on protein interaction and network through alteration of protein structure.
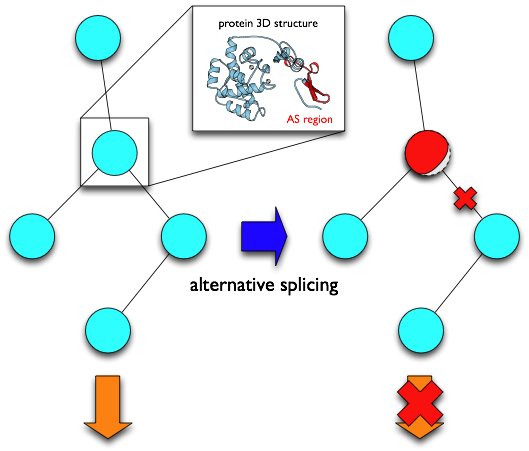
In AS-ALPS, regions of amino acid sequences changed by AS ("AS regions") which we detected with human and mouse transcript sequences in H-InvDB, FANTOM and RefSeq, are linked to information extracted from PDB about residues forming hydrophobic cores and inter-molecular interaction sites. This makes it possible to directly infer whether protein structure and/or interaction are affected by each AS event. In addition, AS-ALPS provides links to a protein network database KEGG, making it easy to know which network and which node in the network can be influenced by AS.
Statistics
Statistics of the database contents in the current version can be viewed from HERE.
Download
Lists of AS genes in ver. 1.6 can be downloaded here.
- Human
- Human including probably artifactual AS genes
- Mouse
- Mouse including probably artifactual AS genes
Publications
If you use AS-ALPS in your work, we request you to cite the following paper and the URL (http://as-alps.nagahama-i-bio.ac.jp/).
Masafumi Shionyu, Akihiro Yamaguchi, Kazuki Shinoda, Ken-ichi Takahashi, and Mitiko Go, 2009. AS-ALPS: a database for analyzing the effects of alternative splicing on protein structure, interaction and network in human and mouse. Nucleic Acids Res. 37(Database issue):D305-D309 [Abstract]
Methodologies to detect AS region and annotate AS regions are described in the following paper.
Kei Yura, Masafumi Shionyu, Kei Hagino, Atsushi Hijikata, Yoshinori Hirashima, Taku Nakahara, Tatsuya Eguchi, Kazuki Shinoda, Akihiro Yamaguchi, Ken-ichi Takahashi, Takeshi Itoh, Tadashi Imanishi, Takashi Gojobori, Mitiko Go, 2006. Alternative splicing in human transcriptome: functional and structural influence on proteins. Gene. 38:63-71 [Abstract]
Acknowledgements
This database was supported by a grant of the Genome Network Project and Grants-in-Aid for Scientific Research (B) from Ministry of Education, Culture, Sports, Science and Technology of Japan.